Merging data
National Data Management Center for Health (NDMC) at EPHI
Data mergining
Why Merging Matters in Public Health
- Combine demographic & clinical data
- Link longitudinal health records
- Merge survey responses with medical data
- Integrate multiple data sources for cohort studies
- The 4 mutating join verbs:
left_join()right_join()inner_join()full_join()
The 2 binding join verbs:
bind_rows()bind_cols()
- The 2 filtering join verbs:
semi_join()anti_join()
- The 3 set operations:
intersect()union()setdiff()
All the joins have this basic syntax:
*_join(x, y, by = NULL, suffix = c(".x", ".y")x =the first (left) tabley =the second (right) tableby =what columns to match on. If you leave this blank, it will match on all columns with the same names in the two tables.suffix =if columns have the same name in the two tables, but you aren’t joining by them, they get a suffix to make them unambiguous.This defaults to “.x” and “.y”, but you can change it to something more meaningful.
Sample Health Datasets
- Patient Demographics (Synthetic)
- Clinical Measurements
left_join(): Preserve Clinic Records
What it does:
Retains all rows from the left (first) table
Adds matching columns from the right (second) table
Fills NA where no match exists
left_joint()
Code
# A tibble: 4 × 7
patient_id age bmi smoking_status visit_date sbp dbp
<chr> <dbl> <dbl> <chr> <date> <dbl> <dbl>
1 P001 35 22.1 former NA NA NA
2 P002 28 26.5 never 2023-01-15 120 80
3 P003 42 29.8 current 2023-02-01 135 85
4 P003 42 29.8 current 2023-03-01 140 90Merging patient registries with lab results
Preserving all patients from primary clinic records
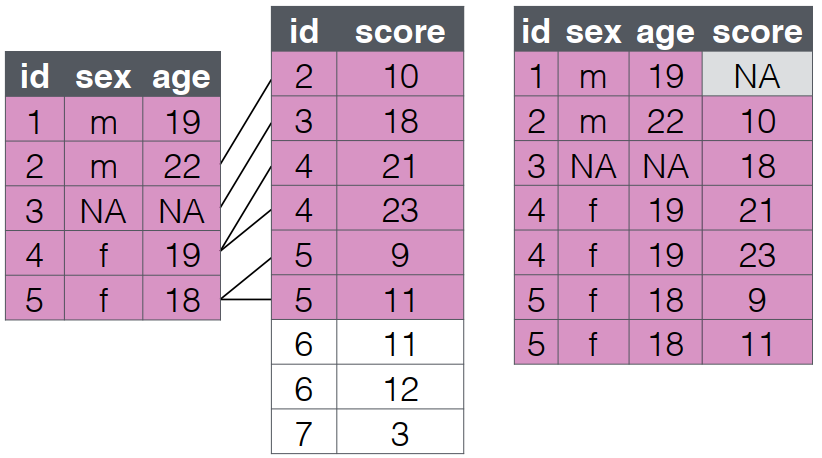
- The order of the
clinic_dataandlab_datatables is different.
# A tibble: 4 × 7
patient_id visit_date sbp dbp age bmi smoking_status
<chr> <date> <dbl> <dbl> <dbl> <dbl> <chr>
1 P002 2023-01-15 120 80 28 26.5 never
2 P003 2023-02-01 135 85 42 29.8 current
3 P003 2023-03-01 140 90 42 29.8 current
4 P005 2023-01-20 128 82 NA NA <NA> 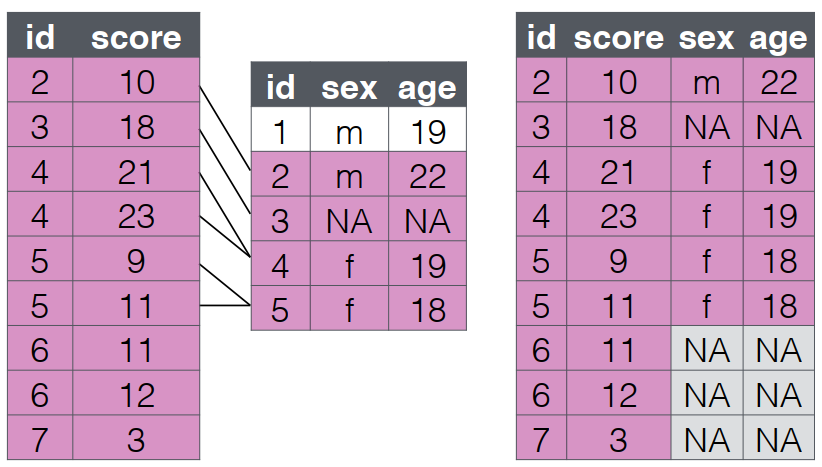
right_join()
- A
right_joinkeeps all the data from the second (right) table and joins anything that matches from the first (left) table.
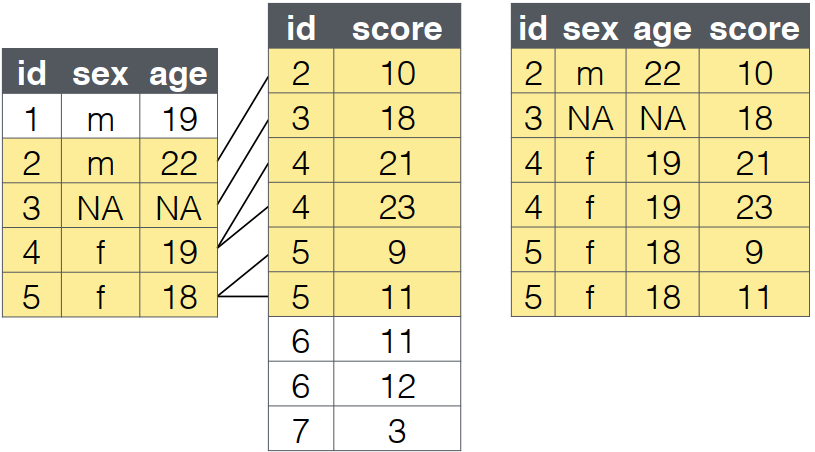
inner_join(): Complete Cases Only
What it does:
- Returns only rows with matches in both tables
- Filters out non-matching records
- An
inner_joinreturns all the rows that have a match in the other table.
| patient_id | age | bmi | smoking_status | visit_date | sbp | dbp |
|---|---|---|---|---|---|---|
| P002 | 28 | 26.5 | never | 2023-01-15 | 120 | 80 |
| P003 | 42 | 29.8 | current | 2023-02-01 | 135 | 85 |
| P003 | 42 | 29.8 | current | 2023-03-01 | 140 | 90 |
- Creating analysis datasets with complete information
- Identifying patients with both survey and clinical data
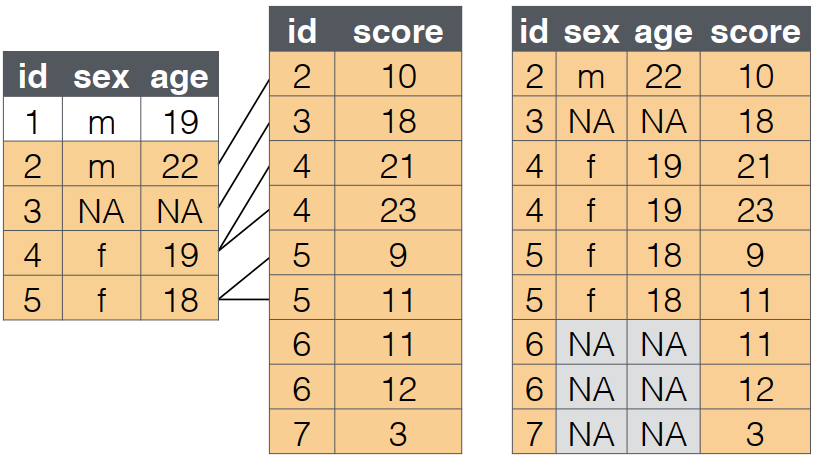
full_join()
What it does:
- A
full_joinlets you join up rows in two tables while keeping all of the information from both tables. - If a row doesn’t have a match in the other table, the other table’s column values are set to
NA.
# A tibble: 6 × 7
patient_id age bmi smoking_status visit_date sbp dbp
<chr> <dbl> <dbl> <chr> <date> <dbl> <dbl>
1 P001 35 22.1 former NA NA NA
2 P002 28 26.5 never 2023-01-15 120 80
3 P003 42 29.8 current 2023-02-01 135 85
4 P003 42 29.8 current 2023-03-01 140 90
5 P004 31 24.3 never NA NA NA
6 P005 NA NA <NA> 2023-01-20 128 82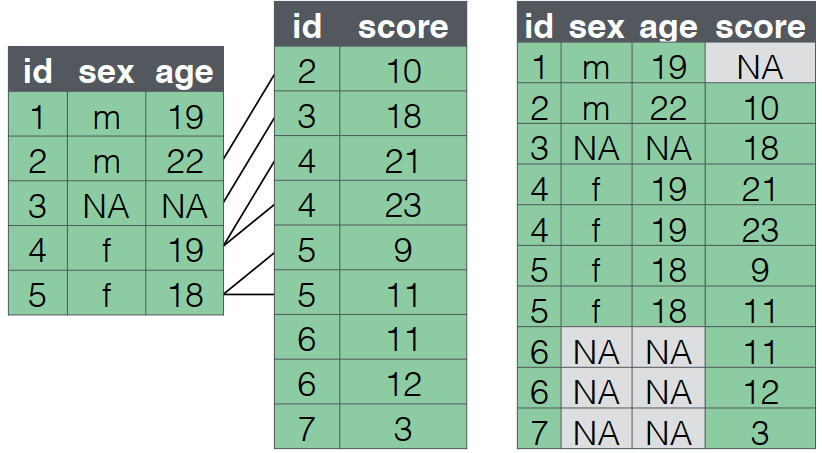
semi_join()
- A
semi_joinfilters left table to rows with matches in right table - Keeps only left table columns
# A tibble: 2 × 4
patient_id age bmi smoking_status
<chr> <dbl> <dbl> <chr>
1 P002 28 26.5 never
2 P003 42 29.8 current 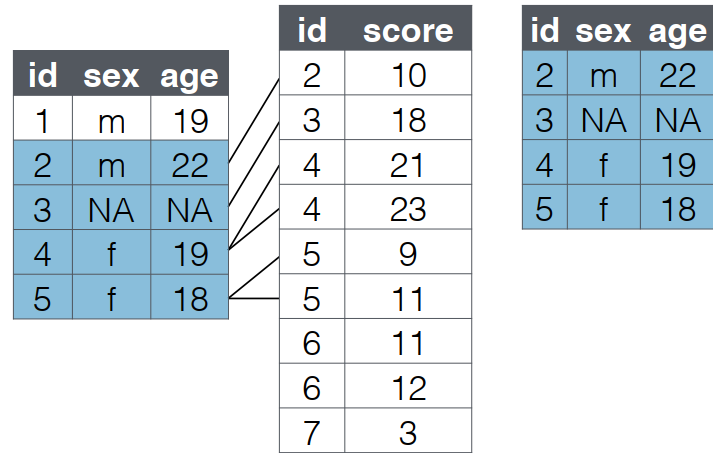
- Here in this data case: Find patients with lab results
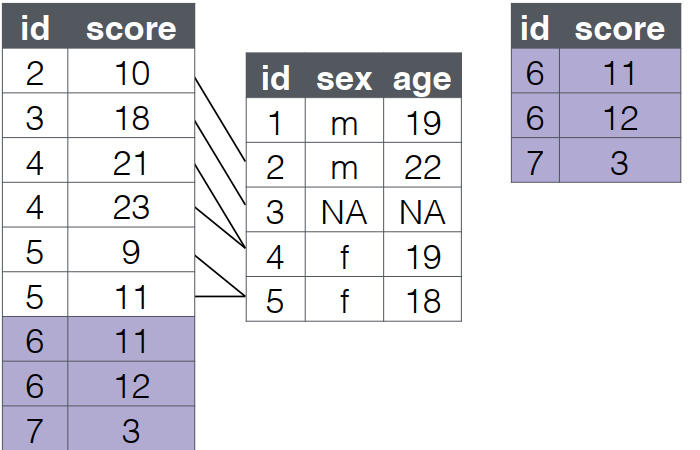
anti_join(): Identify Missing Data
- A
anti_join()return all rows from the left table where there are not matching values in the right table, keeping just columns from the left table.
- Order matters in an anti_join().
| patient_id | visit_date | sbp | dbp |
|---|---|---|---|
| P005 | 2023-01-20 | 128 | 82 |
:::::::
Find patients needing follow-up lab tests
Real-world Example: Birth Weight Analysis Birth Weight Data (MASS::birthwt)
Postnatal Follow-up Data (Synthetic)
- full_join(): Comprehensive Birth Cohort
- Combine prenatal and postnatal data for longitudinal analysis
Code
| patient_id | low | smoke | infant_weight_6mo |
|---|---|---|---|
| M12 | Normal | Non-smoker | 7.2 |
| M15 | Normal | Smoker | 6.8 |
| M42 | Normal | Non-smoker | 7.5 |
| M88 | Normal | Non-smoker | 6.9 |
- semi_join(): Smoking Mothers with Follow-up
- Identify smoking mothers in follow-up program
bind_rows()
- You can combine the rows of two tables with
bind_rows. - Here we’ll add subject data for subjects 6-9 and bind that to the original subject table.
Code
# A tibble: 6 × 4
patient_id age bmi smoking_status
<chr> <dbl> <dbl> <chr>
1 P001 35 22.1 former
2 P002 28 26.5 never
3 P003 42 29.8 current
4 P004 31 24.3 never
5 P4 55 NA Never
6 P5 38 NA Current 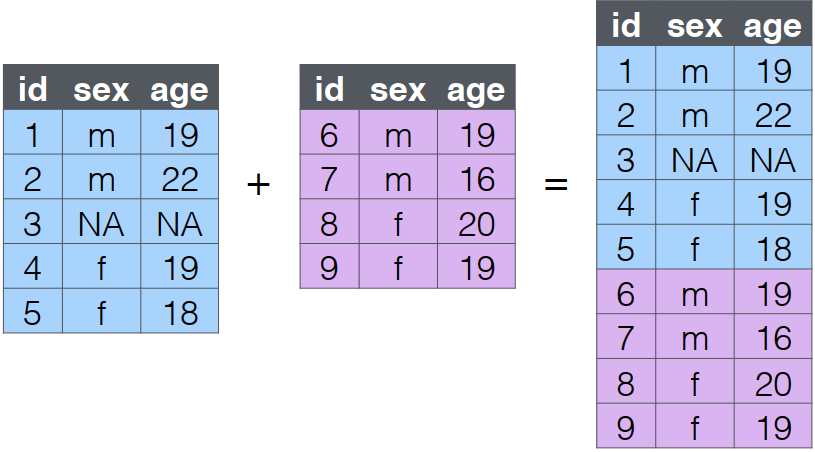
- The columns just have to have the same names, they don’t have to be in the same order.
Any columns that differ between the two tables will just have
NAvalues for entries from the other table.If a row is duplicated between the two tables (like id 5 below), the row will also be duplicated in the resulting table.
